---
title: "Calculate SPEI"
subtitle: "Standardized Precipitation Evapotranspiration Index - Complete Tutorial"
author: "Benny Istanto"
date: last-modified
execute:
eval: true
echo: true
warning: false
cache: true
format:
html:
code-fold: show
code-tools: true
---
## Overview
This tutorial demonstrates how to calculate the **Standardized Precipitation Evapotranspiration Index (SPEI)** using TerraClimate data for Bali, Indonesia (1958-2024).
**What is SPEI?**
- Extension of SPI that includes temperature/evapotranspiration
- Developed by Vicente-Serrano et al. (2010)
- Uses water balance (P - PET) instead of precipitation alone
- More sensitive to temperature changes and global warming
- Better for agricultural and ecological drought assessment
**What you'll learn:**
1. Load and explore precipitation, temperature, and PET data
2. Choose between pre-computed PET or calculating from temperature
3. Visualize input climate data (time series and spatial patterns)
4. Calculate SPEI-12 and visualize results
5. Compare SPI vs SPEI to see the effect of temperature
6. Calculate multi-scale SPEI (1, 3, 6, 12 months)
7. Use different probability distributions (Gamma, Pearson III, Log-Logistic)
8. Spatial analysis of drought conditions
9. Drought area percentage comparison (SPI vs SPEI)
10. Save and reuse fitting parameters
11. Save results to NetCDF
## Why SPEI?
**SPEI** extends SPI by incorporating **potential evapotranspiration (PET)**:
- **SPI** = Precipitation only
- **SPEI** = Precipitation - PET (water balance)
SPEI is more sensitive to temperature increases and climate change impacts.
## 1. Setup and Imports
```{python}
import sys
import os
from pathlib import Path
# Get the notebook directory and find the repository root
notebook_dir = Path.cwd()
repo_root = notebook_dir
while not (repo_root / 'src').exists() and repo_root != repo_root.parent:
repo_root = repo_root.parent
# Add src to path
src_path = str(repo_root / 'src')
if src_path not in sys.path:
sys.path.insert(0, src_path)
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
from datetime import datetime
# Import SPEI and SPI functions
from indices import (
spi,
spei,
spei_multi_scale,
save_fitting_params,
load_fitting_params,
save_index_to_netcdf,
get_drought_area_percentage
)
from utils import calculate_pet
from visualization import plot_index
from config import Periodicity, DISTRIBUTION_DISPLAY_NAMES
# Plotting settings
plt.rcParams['figure.figsize'] = (12, 6)
plt.rcParams['font.size'] = 10
# Create output directories
output_plots = repo_root / 'output' / 'plots'
output_netcdf = repo_root / 'output' / 'netcdf'
output_plots.mkdir(parents=True, exist_ok=True)
output_netcdf.mkdir(parents=True, exist_ok=True)
print("All imports successful!")
print(f"Repository root: {repo_root}")
```
## 2. Configuration
Choose your PET input method and set calculation parameters.
**You have THREE options for SPEI calculation:**
- **Option A**: Use pre-computed PET (recommended if available) --- faster, uses existing PET from TerraClimate
- **Option B**: Calculate PET from temperature (Thornthwaite) --- requires only mean temperature data
- **Option C**: Calculate PET using Hargreaves --- requires Tmin/Tmax data, better for arid regions
```{python}
# ========================================
# USER CONFIGURATION
# ========================================
# Choose PET input method
use_precomputed_pet = False # Set True for pre-computed PET, False to calculate from temperature
# File paths
precip_file = repo_root / 'input' / 'terraclimate_bali_ppt_1958_2024.nc'
pet_file = repo_root / 'input' / 'terraclimate_bali_pet_1958_2024.nc'
temp_file = repo_root / 'input' / 'terraclimate_bali_tmean_1958_2024.nc'
# Variable names in your NetCDF files
precip_var = 'ppt'
pet_var = 'pet'
temp_var = 'tmean'
# Calibration period (WMO standard: 1991-2020)
calib_start = 1991
calib_end = 2020
data_start_year = 1958
print("Configuration:")
print(f" PET method: {'Pre-computed' if use_precomputed_pet else 'Calculate from temperature'}")
print(f" Calibration period: {calib_start}-{calib_end}")
print(f" Output directory: {output_netcdf}")
```
::: {.callout-note}
### PET Calculation Methods
The `spei()` function accepts:
- `pet=` parameter for pre-computed PET values (e.g., from TerraClimate)
- `temperature=` + `latitude=` to auto-calculate PET using Thornthwaite method (default)
- `temperature=` + `temp_min=` + `temp_max=` + `latitude=` + `pet_method='hargreaves'` for Hargreaves-Samani method
**Thornthwaite** is simpler (requires only mean temperature), while **Hargreaves** performs better in arid/semi-arid regions and correlates better with physically-based PET like Penman-Monteith.
:::
## 3. Load Climate Data
```{python}
# Load precipitation data
print("Loading precipitation data...")
ds_precip = xr.open_dataset(precip_file)
precip = ds_precip[precip_var]
print("Precipitation data loaded")
print(f" Shape: {precip.shape}")
print(f" Time range: {precip.time.min().values} to {precip.time.max().values}")
print(f" Spatial extent: lat [{precip.lat.min().values:.2f}, {precip.lat.max().values:.2f}], "
f"lon [{precip.lon.min().values:.2f}, {precip.lon.max().values:.2f}]")
print(f" Mean: {precip.mean().values:.1f} mm/month")
print(f" Range: [{precip.min().values:.1f}, {precip.max().values:.1f}] mm/month")
```
```{python}
# Load PET data (Option A) or Temperature data (Option B)
if use_precomputed_pet:
print("\nLoading pre-computed PET data...")
ds_pet = xr.open_dataset(pet_file)
pet = ds_pet[pet_var]
print("PET data loaded")
print(f" Shape: {pet.shape}")
print(f" Mean: {pet.mean().values:.1f} mm/month")
print(f" Range: [{pet.min().values:.1f}, {pet.max().values:.1f}] mm/month")
temperature = None
else:
print("\nLoading temperature data for PET calculation...")
ds_temp = xr.open_dataset(temp_file)
temperature = ds_temp[temp_var]
print("Temperature data loaded")
print(f" Shape: {temperature.shape}")
print(f" Mean: {temperature.mean().values:.1f} deg C")
print(f" Range: [{temperature.min().values:.1f}, {temperature.max().values:.1f}] deg C")
pet = None
```
## 4. Visualize Climate Data
Examine the climate data at a sample location before calculating SPEI.
```{python}
# Select a sample location (center of domain)
sample_lat_idx = len(precip.lat) // 2
sample_lon_idx = len(precip.lon) // 2
sample_lat = precip.lat[sample_lat_idx].values
sample_lon = precip.lon[sample_lon_idx].values
print(f"Sample location: lat={sample_lat:.3f}, lon={sample_lon:.3f}")
```
```{python}
if use_precomputed_pet:
# Show Precipitation, PET, and Water Balance
fig, axes = plt.subplots(3, 1, figsize=(14, 10))
# Precipitation
precip[:, sample_lat_idx, sample_lon_idx].plot(ax=axes[0], linewidth=0.8, color='steelblue')
axes[0].set_title(f'Precipitation (lat={sample_lat:.3f}, lon={sample_lon:.3f})')
axes[0].set_ylabel('Precipitation (mm/month)')
axes[0].grid(True, alpha=0.3)
# PET
pet[:, sample_lat_idx, sample_lon_idx].plot(ax=axes[1], linewidth=0.8, color='orangered')
axes[1].set_title('Potential Evapotranspiration (PET)')
axes[1].set_ylabel('PET (mm/month)')
axes[1].grid(True, alpha=0.3)
# Water Balance (P - PET)
water_balance = precip - pet
wb = water_balance[:, sample_lat_idx, sample_lon_idx]
wb.plot(ax=axes[2], linewidth=0.8, color='darkgreen')
axes[2].axhline(y=0, color='k', linestyle='-', linewidth=0.8, alpha=0.3)
axes[2].fill_between(wb.time, 0, wb, where=(wb >= 0), alpha=0.3, color='blue', label='Surplus')
axes[2].fill_between(wb.time, 0, wb, where=(wb < 0), alpha=0.3, color='red', label='Deficit')
axes[2].set_title('Water Balance (P - PET)')
axes[2].set_ylabel('P - PET (mm/month)')
axes[2].grid(True, alpha=0.3)
axes[2].legend()
plt.tight_layout()
plt.show()
# Statistics
print(f"\nWater Balance Statistics:")
print(f" Mean P: {precip.mean().values:.1f} mm/month")
print(f" Mean PET: {pet.mean().values:.1f} mm/month")
print(f" Mean (P-PET): {water_balance.mean().values:.1f} mm/month")
deficit_frac = float((water_balance < 0).sum() / water_balance.size)
print(f" Deficit months: {int((water_balance < 0).sum())} ({100*deficit_frac:.1f}%)")
else:
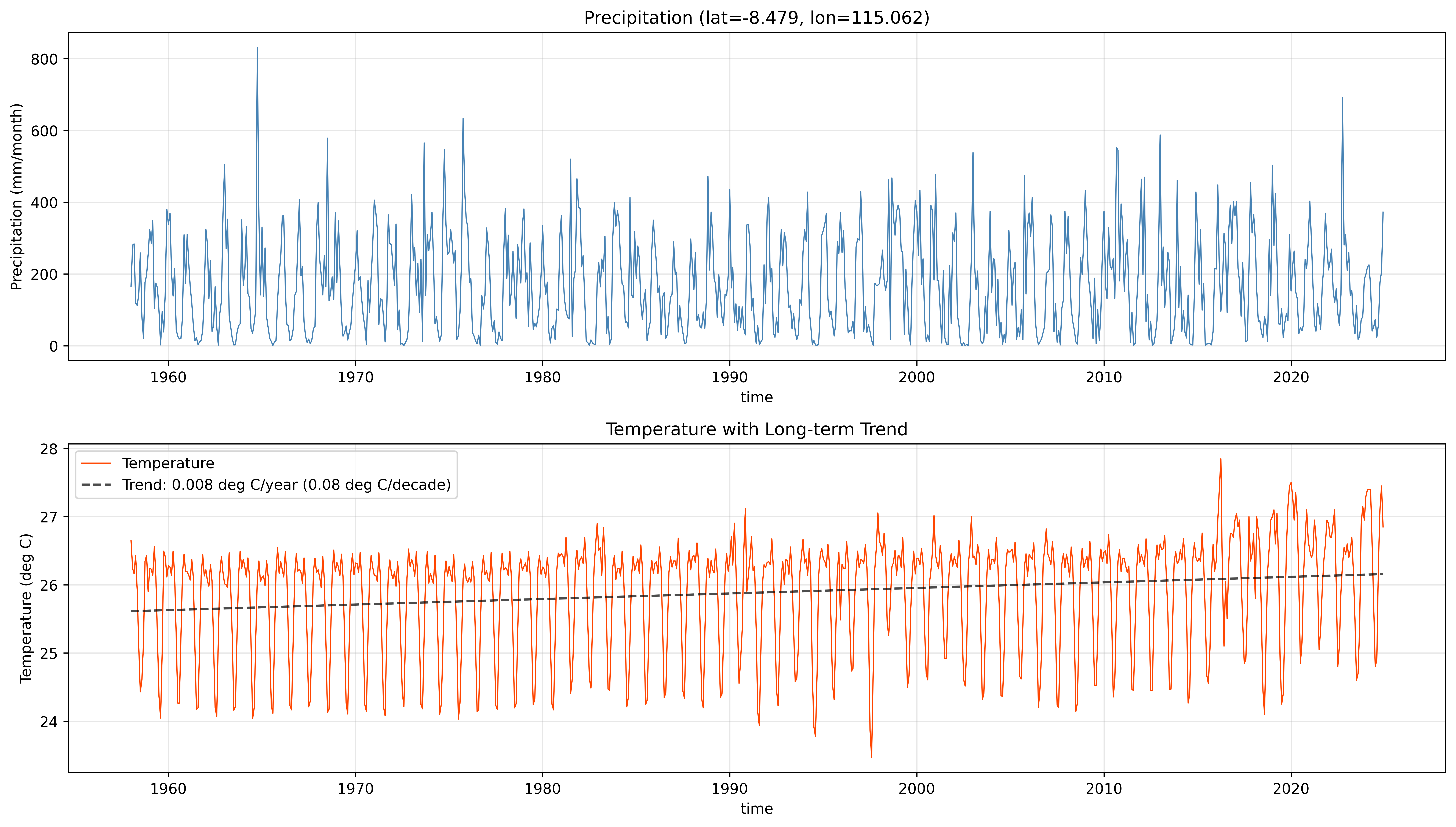
# Show Precipitation and Temperature
fig, axes = plt.subplots(2, 1, figsize=(14, 8))
# Precipitation
precip[:, sample_lat_idx, sample_lon_idx].plot(ax=axes[0], linewidth=0.8, color='steelblue')
axes[0].set_title(f'Precipitation (lat={sample_lat:.3f}, lon={sample_lon:.3f})')
axes[0].set_ylabel('Precipitation (mm/month)')
axes[0].grid(True, alpha=0.3)
# Temperature with trend
temp_series = temperature[:, sample_lat_idx, sample_lon_idx]
temp_series.plot(ax=axes[1], linewidth=0.8, color='orangered', label='Temperature')
# Add trend line
x = np.arange(len(temp_series))
valid_mask = ~np.isnan(temp_series.values)
if valid_mask.sum() > 0:
z = np.polyfit(x[valid_mask], temp_series.values[valid_mask], 1)
p = np.poly1d(z)
axes[1].plot(temp_series.time, p(x), 'k--', linewidth=1.5, alpha=0.7,
label=f'Trend: {z[0]*12:.3f} deg C/year ({z[0]*120:.2f} deg C/decade)')
axes[1].set_title('Temperature with Long-term Trend')
axes[1].set_ylabel('Temperature (deg C)')
axes[1].grid(True, alpha=0.3)
axes[1].legend()
plt.tight_layout()
plt.show()
print(f"\nTemperature Statistics:")
print(f" Mean: {temperature.mean().values:.1f} deg C")
print(f" Range: [{temperature.min().values:.1f}, {temperature.max().values:.1f}] deg C")
```
When using temperature, the trend line shows how temperatures have changed over the record. Warming trends increase PET, which affects the water balance and makes SPEI more negative relative to SPI.
::: {.callout-tip}
### Distribution Selection for SPEI
By default, SPEI uses the **Gamma** distribution. For SPEI, the **Pearson III** or **Log-Logistic** distributions are also available and may better handle the water balance data (P - PET), which can be negative:
```python
# SPEI with Pearson III
spei_6_p3 = spei(precip, pet=pet_precomputed, scale=6, distribution='pearson3')
# SPEI with Log-Logistic (as in original R SPEI package)
spei_6_ll = spei(precip, pet=pet_precomputed, scale=6, distribution='log_logistic')
```
:::
## 5. Calculate SPEI-12
Calculate SPEI at 12-month timescale for long-term water resources monitoring.
```{python}
print("Calculating SPEI-12 (Gamma distribution)...")
print(f" Method: {'Using pre-computed PET' if use_precomputed_pet else 'Calculating PET from temperature'}")
if use_precomputed_pet:
spei_12, params = spei(
precip=precip,
pet=pet,
scale=12,
periodicity='monthly',
calibration_start_year=calib_start,
calibration_end_year=calib_end,
distribution='gamma', # Default; also supports 'pearson3', 'log_logistic'
return_params=True
)
else:
spei_12, params = spei(
precip=precip,
temperature=temperature,
latitude=precip.lat.values,
scale=12,
periodicity='monthly',
calibration_start_year=calib_start,
calibration_end_year=calib_end,
distribution='gamma', # Default; also supports 'pearson3', 'log_logistic'
return_params=True
)
print("SPEI-12 calculation complete!")
print(f" Output shape: {spei_12.shape}")
print(f" Valid range: [{float(np.nanmin(spei_12.values)):.2f}, {float(np.nanmax(spei_12.values)):.2f}]")
print(f" Mean: {float(np.nanmean(spei_12.values)):.3f}")
print(f" Std: {float(np.nanstd(spei_12.values)):.3f}")
```
### Visualize SPEI-12 Results
```{python}
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(14, 12))
# Time series at sample point
spei_series = spei_12[:, sample_lat_idx, sample_lon_idx]
spei_series.plot(ax=ax1, linewidth=0.8, color='darkgreen')
ax1.axhline(y=0, color='k', linestyle='-', linewidth=0.8, alpha=0.3)
ax1.axhline(y=-1.0, color='orange', linestyle='--', linewidth=0.8, alpha=0.5, label='Moderate dry/wet')
ax1.axhline(y=-2.0, color='red', linestyle='--', linewidth=0.8, alpha=0.5, label='Severe dry/wet')
ax1.axhline(y=1.0, color='orange', linestyle='--', linewidth=0.8, alpha=0.5)
ax1.axhline(y=2.0, color='red', linestyle='--', linewidth=0.8, alpha=0.5)
ax1.fill_between(spei_12.time, -10, 0, alpha=0.1, color='red', label='Dry')
ax1.fill_between(spei_12.time, 0, 10, alpha=0.1, color='blue', label='Wet')
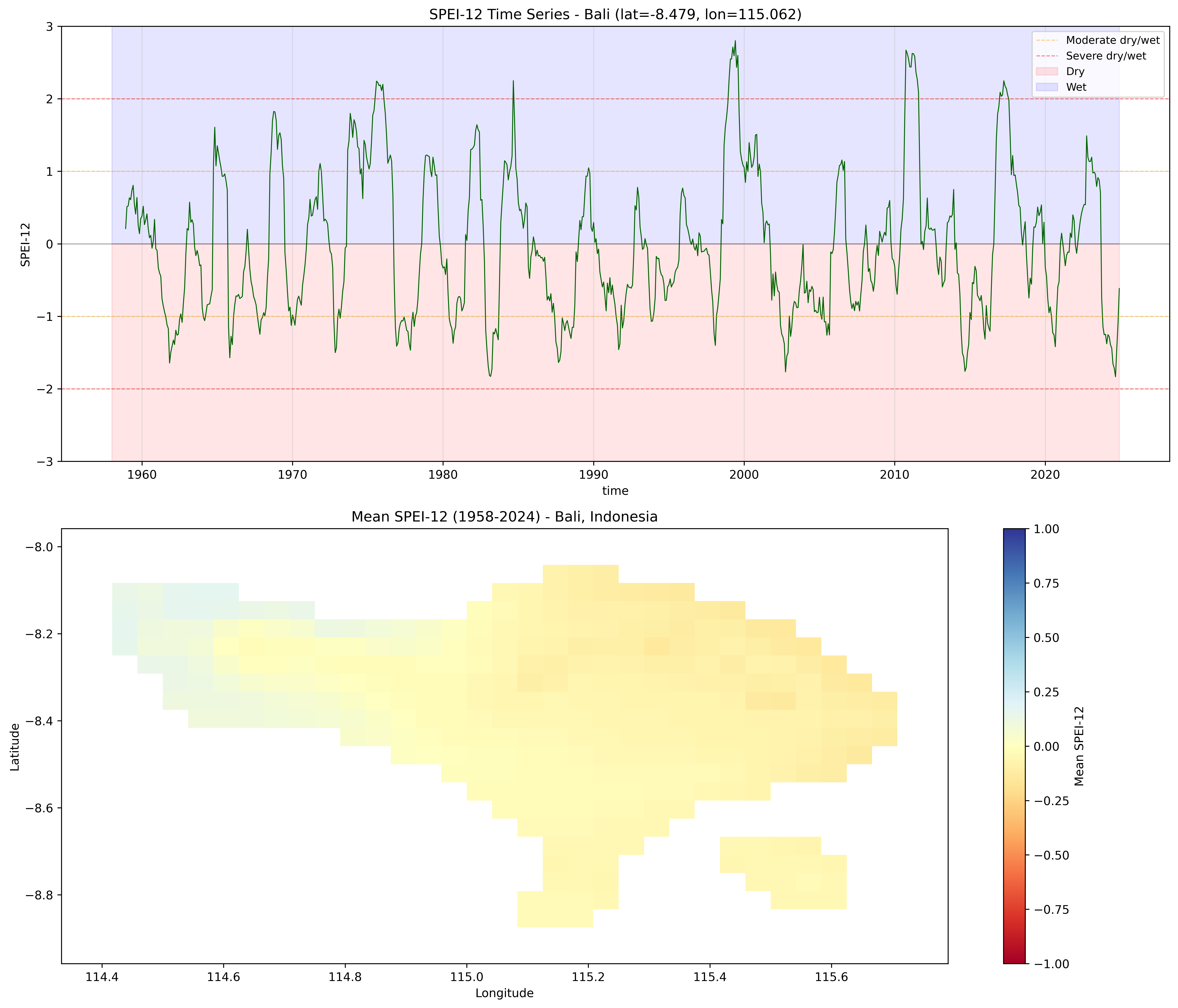
ax1.set_title(f'SPEI-12 Time Series - Bali (lat={sample_lat:.3f}, lon={sample_lon:.3f})')
ax1.set_ylabel('SPEI-12')
ax1.set_ylim(-3, 3)
ax1.grid(True, alpha=0.3)
ax1.legend(loc='upper right', fontsize=9)
# Spatial pattern (mean over time)
spei_12.mean(dim='time').plot(ax=ax2, cmap='RdYlBu', vmin=-1, vmax=1,
cbar_kwargs={'label': 'Mean SPEI-12'})
ax2.set_title('Mean SPEI-12 (1958-2024) - Bali, Indonesia')
ax2.set_xlabel('Longitude')
ax2.set_ylabel('Latitude')
plt.tight_layout()
plt.show()
```
The top panel shows the SPEI-12 time series with threshold lines for moderate (+-1.0) and severe (+-2.0) conditions. The bottom panel shows the spatial mean SPEI-12 across Bali.
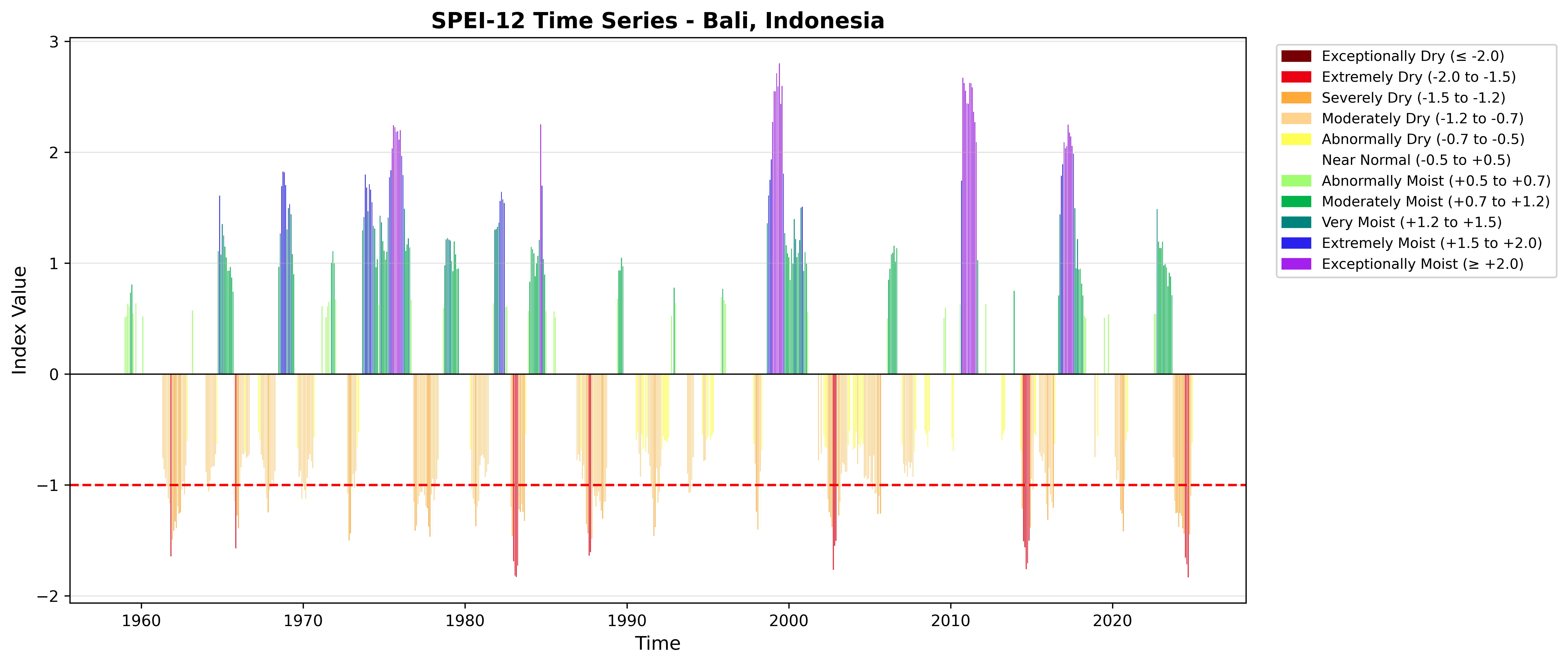
### SPEI-12 with WMO Color Scheme
```{python}
spei_point = spei_12.isel(lat=sample_lat_idx, lon=sample_lon_idx)
fig = plot_index(
spei_point,
threshold=-1.0,
title='SPEI-12 Time Series - Bali, Indonesia'
)
plt.tight_layout()
plt.show()
```
## 6. Compare SPI vs SPEI
Calculate SPI-12 and compare with SPEI-12 to see the effect of including temperature/PET.
```{python}
print("Calculating SPI-12 for comparison...")
spi_12 = spi(
precip=precip,
scale=12,
periodicity='monthly',
calibration_start_year=calib_start,
calibration_end_year=calib_end
)
print("SPI-12 calculation complete!")
```
```{python}
# Compare SPI and SPEI at sample location
fig, axes = plt.subplots(3, 1, figsize=(14, 11))
# SPI
spi_series = spi_12[:, sample_lat_idx, sample_lon_idx]
spi_series.plot(ax=axes[0], linewidth=0.8, color='steelblue', label='SPI-12')
axes[0].axhline(y=0, color='k', linestyle='-', linewidth=0.8, alpha=0.3)
axes[0].axhline(y=-1.0, color='orange', linestyle='--', linewidth=0.6, alpha=0.4, label='Moderate drought')
axes[0].axhline(y=-2.0, color='red', linestyle='--', linewidth=0.6, alpha=0.4, label='Severe drought')
axes[0].fill_between(spi_series.time, -10, 0, alpha=0.1, color='red')
axes[0].fill_between(spi_series.time, 0, 10, alpha=0.1, color='blue')
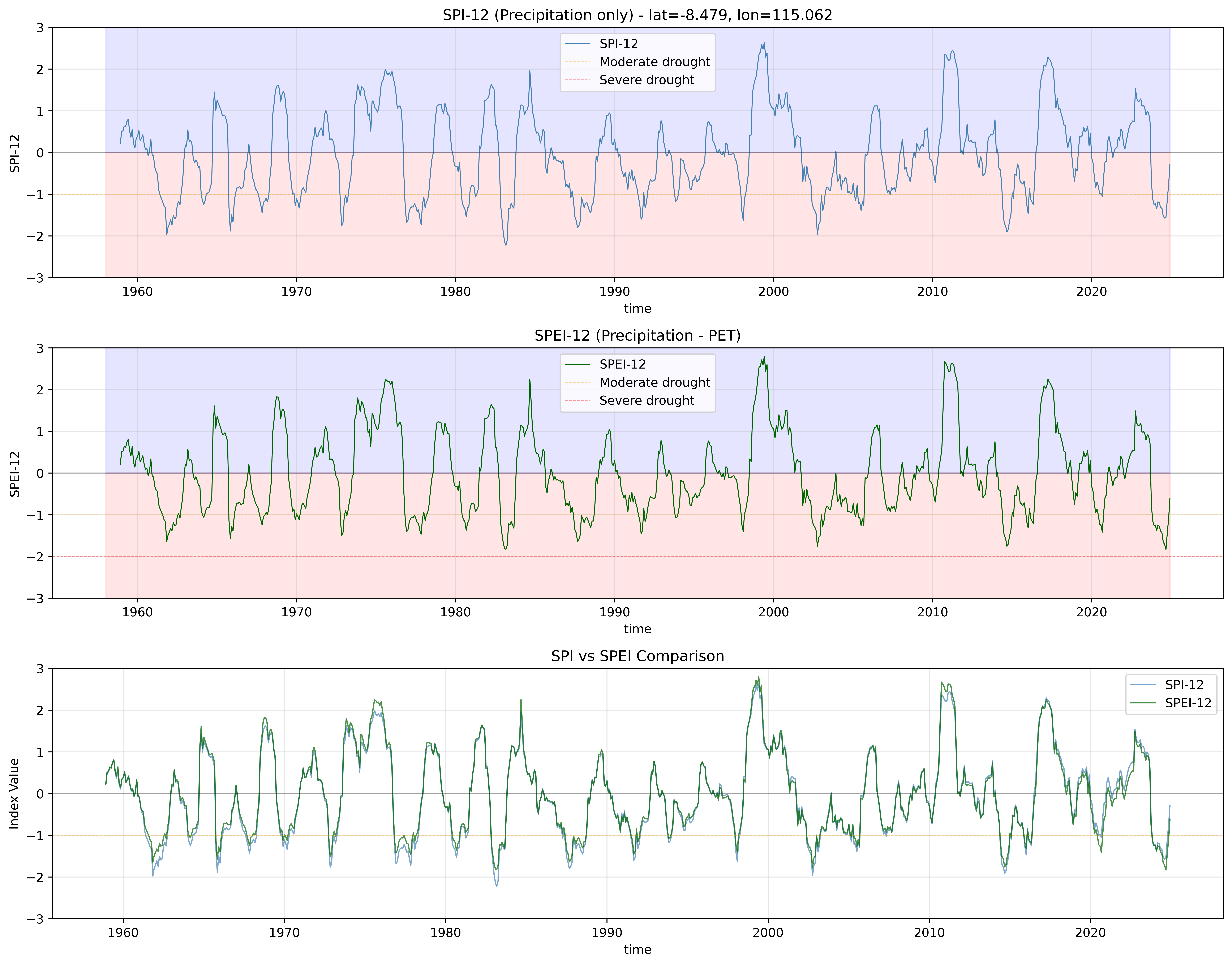
axes[0].set_title(f'SPI-12 (Precipitation only) - lat={sample_lat:.3f}, lon={sample_lon:.3f}')
axes[0].set_ylabel('SPI-12')
axes[0].set_ylim(-3, 3)
axes[0].grid(True, alpha=0.3)
axes[0].legend()
# SPEI
spei_series = spei_12[:, sample_lat_idx, sample_lon_idx]
spei_series.plot(ax=axes[1], linewidth=0.8, color='darkgreen', label='SPEI-12')
axes[1].axhline(y=0, color='k', linestyle='-', linewidth=0.8, alpha=0.3)
axes[1].axhline(y=-1.0, color='orange', linestyle='--', linewidth=0.6, alpha=0.4, label='Moderate drought')
axes[1].axhline(y=-2.0, color='red', linestyle='--', linewidth=0.6, alpha=0.4, label='Severe drought')
axes[1].fill_between(spei_series.time, -10, 0, alpha=0.1, color='red')
axes[1].fill_between(spei_series.time, 0, 10, alpha=0.1, color='blue')
axes[1].set_title('SPEI-12 (Precipitation - PET)')
axes[1].set_ylabel('SPEI-12')
axes[1].set_ylim(-3, 3)
axes[1].grid(True, alpha=0.3)
axes[1].legend()
# Overlay comparison
spi_series.plot(ax=axes[2], linewidth=1.0, color='steelblue', label='SPI-12', alpha=0.7)
spei_series.plot(ax=axes[2], linewidth=1.0, color='darkgreen', label='SPEI-12', alpha=0.7)
axes[2].axhline(y=0, color='k', linestyle='-', linewidth=0.8, alpha=0.3)
axes[2].axhline(y=-1.0, color='orange', linestyle='--', linewidth=0.6, alpha=0.4)
axes[2].set_title('SPI vs SPEI Comparison')
axes[2].set_ylabel('Index Value')
axes[2].set_ylim(-3, 3)
axes[2].grid(True, alpha=0.3)
axes[2].legend()
plt.tight_layout()
plt.show()
# Calculate statistics
valid_mask = ~(np.isnan(spi_series.values) | np.isnan(spei_series.values))
if valid_mask.sum() > 0:
correlation = np.corrcoef(spi_series.values[valid_mask], spei_series.values[valid_mask])[0, 1]
print(f"\nCorrelation between SPI and SPEI: {correlation:.3f}")
# Count drought months
spi_drought = int((spi_series <= -1.0).sum().values)
spei_drought = int((spei_series <= -1.0).sum().values)
total_months = len(spi_series)
print(f"\nDrought months (index <= -1.0):")
print(f" SPI-12: {spi_drought} months ({100*spi_drought/total_months:.1f}%)")
print(f" SPEI-12: {spei_drought} months ({100*spei_drought/total_months:.1f}%)")
print(f" Difference: {spei_drought - spi_drought} months ({100*(spei_drought - spi_drought)/total_months:.1f}%)")
```
::: {.callout-important}
### SPI vs SPEI: Key Differences
**SPI (Precipitation only):**
- Simple, requires only precipitation data
- Good for meteorological drought
- Does not account for temperature/evaporation
- May miss agricultural drought in warming climates
**SPEI (Precipitation - PET):**
- Accounts for temperature and evaporation
- Better for agricultural and ecological drought
- More sensitive to climate change
- Requires temperature data or PET
**When to use SPEI:**
- Agricultural applications (crop monitoring)
- Vegetation/ecosystem monitoring
- Climate change impact studies
- Regions with significant temperature trends
- Forest fire risk assessment
:::
## 7. Multi-Scale SPEI Analysis
Different time scales capture different types of drought:
- **SPEI-1**: Short-term meteorological
- **SPEI-3**: Seasonal agricultural
- **SPEI-6**: Medium-term hydrological
- **SPEI-12**: Long-term water resources
```{python}
print("Calculating SPEI for scales: 1, 3, 6, 12 months...")
if use_precomputed_pet:
spei_multi = spei_multi_scale(
precip=precip,
pet=pet,
scales=[1, 3, 6, 12],
periodicity='monthly',
calibration_start_year=calib_start,
calibration_end_year=calib_end
)
else:
spei_multi = spei_multi_scale(
precip=precip,
temperature=temperature,
latitude=precip.lat.values,
scales=[1, 3, 6, 12],
periodicity='monthly',
calibration_start_year=calib_start,
calibration_end_year=calib_end
)
print("Multi-scale SPEI calculation complete!")
print(f" Variables: {list(spei_multi.data_vars)}")
```
### Compare Different Time Scales
```{python}
fig, axes = plt.subplots(4, 1, figsize=(14, 13), sharex=True)
scales = [1, 3, 6, 12]
colors = ['lightgreen', 'mediumseagreen', 'seagreen', 'darkgreen']
for i, (scale, color) in enumerate(zip(scales, colors)):
var_name = f'spei_gamma_{scale}_month'
spei_data = spei_multi[var_name]
spei_data[:, sample_lat_idx, sample_lon_idx].plot(
ax=axes[i],
linewidth=0.8,
color=color
)
axes[i].axhline(y=0, color='k', linestyle='-', linewidth=0.8, alpha=0.3)
axes[i].axhline(y=-1.0, color='orange', linestyle='--', linewidth=0.6, alpha=0.4)
axes[i].axhline(y=-2.0, color='red', linestyle='--', linewidth=0.6, alpha=0.4)
axes[i].axhline(y=1.0, color='orange', linestyle='--', linewidth=0.6, alpha=0.4)
axes[i].axhline(y=2.0, color='red', linestyle='--', linewidth=0.6, alpha=0.4)
axes[i].fill_between(spei_data.time, -10, 0, alpha=0.1, color='red')
axes[i].fill_between(spei_data.time, 0, 10, alpha=0.1, color='blue')
axes[i].set_title(f'SPEI-{scale}')
axes[i].set_ylabel(f'SPEI-{scale}')
axes[i].set_ylim(-3, 3)
axes[i].grid(True, alpha=0.3)
if i < 3:
axes[i].set_xlabel('')
plt.suptitle(f'SPEI Multi-Scale Comparison (lat={sample_lat:.3f}, lon={sample_lon:.3f})',
fontsize=14, y=0.995)
plt.tight_layout()
plt.show()
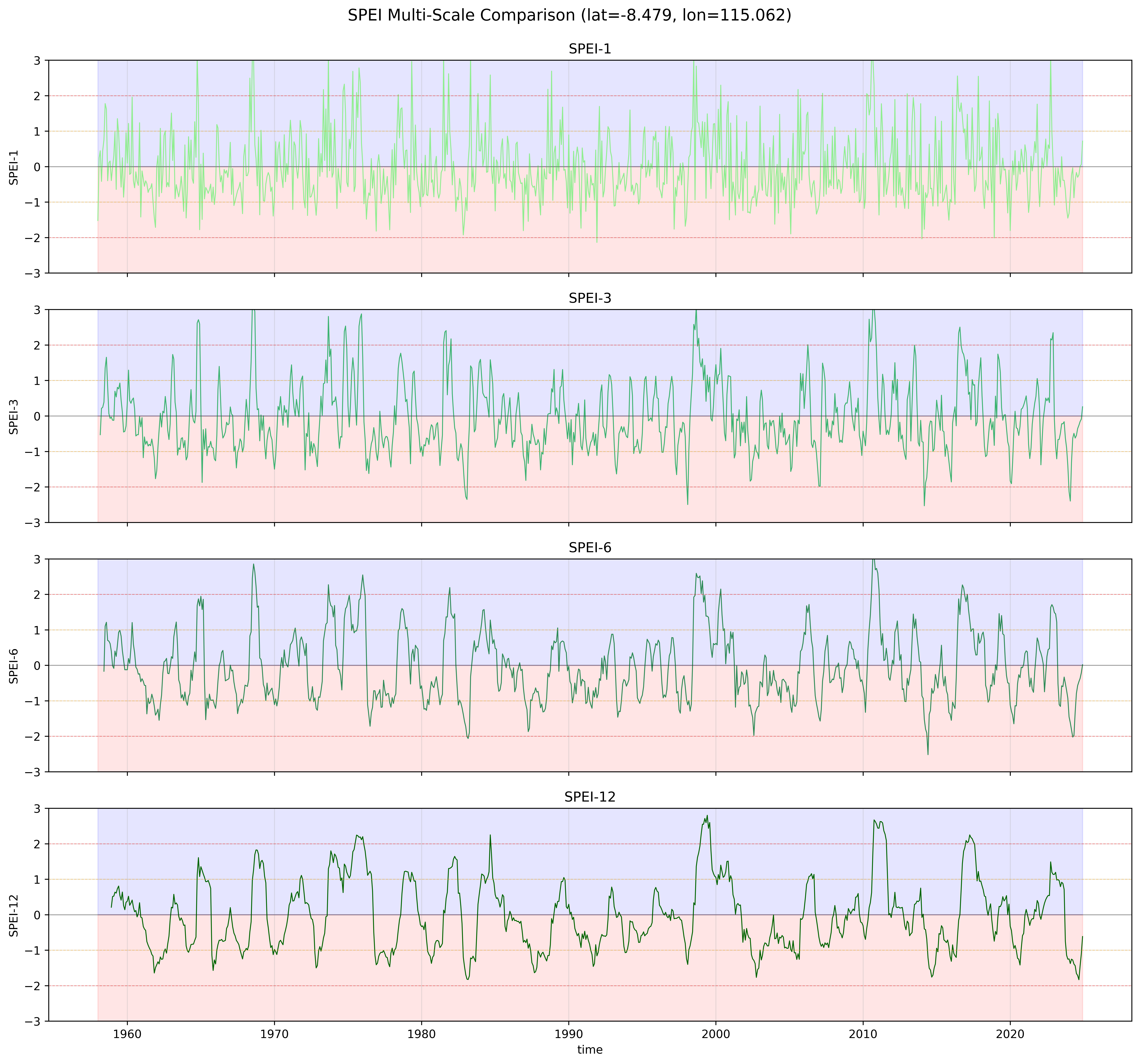
print("\nObservations:")
print(" - SPEI-1 shows high frequency variability (month-to-month)")
print(" - SPEI-12 shows smoother, longer-term trends")
print(" - Extreme events persist longer at longer time scales")
```
::: {.callout-note}
### Time Scale Interpretation
- **SPEI-1:** Month-to-month meteorological anomalies
- **SPEI-3:** Seasonal agricultural drought --- crop water stress
- **SPEI-6:** Medium-term agricultural/hydrological drought
- **SPEI-12:** Long-term hydrological drought --- water resources, reservoir management
:::
## 8. Multi-Distribution Support
The `spei()` function supports three probability distributions:
| Distribution | Parameter | Fitting Method | Notes |
|:------------|:----------|:--------------|:------|
| **Gamma** | `'gamma'` | Method of Moments | Default, fast and robust |
| **Pearson III** | `'pearson3'` | Method of Moments | Better for skewed water balance data |
| **Log-Logistic** | `'log_logistic'` | Maximum Likelihood (MLE) | Used in the original R SPEI package |
```{python}
# Calculate SPEI-6 with each distribution for comparison
print("Comparing SPEI-6 across distributions...")
spei6_results = {}
for dist in ['gamma', 'pearson3', 'log_logistic']:
dist_name = DISTRIBUTION_DISPLAY_NAMES[dist]
if use_precomputed_pet:
spei_6 = spei(
precip=precip, pet=pet, scale=6,
periodicity='monthly',
calibration_start_year=calib_start,
calibration_end_year=calib_end,
distribution=dist
)
else:
spei_6 = spei(
precip=precip, temperature=temperature,
latitude=precip.lat.values, scale=6,
periodicity='monthly',
calibration_start_year=calib_start,
calibration_end_year=calib_end,
distribution=dist
)
spei6_results[dist] = spei_6
print(f" {dist_name}: range [{float(spei_6.min()):.2f}, {float(spei_6.max()):.2f}], mean={float(spei_6.mean()):.3f}")
```
### Compare Distributions
```{python}
fig, axes = plt.subplots(3, 1, figsize=(14, 10), sharex=True, sharey=True)
for ax, dist in zip(axes, ['gamma', 'pearson3', 'log_logistic']):
dist_name = DISTRIBUTION_DISPLAY_NAMES[dist]
data = spei6_results[dist][:, sample_lat_idx, sample_lon_idx].values
colors = np.where(data >= 0, '#2166ac', '#b2182b')
ax.bar(range(len(data)), data, color=colors, width=1.0, edgecolor='none')
ax.axhline(y=0, color='black', linewidth=0.5)
ax.set_ylabel('SPEI-6')
ax.set_title(f'SPEI-6 - {dist_name}')
ax.set_ylim(-3.5, 3.5)
axes[-1].set_xlabel('Time Step')
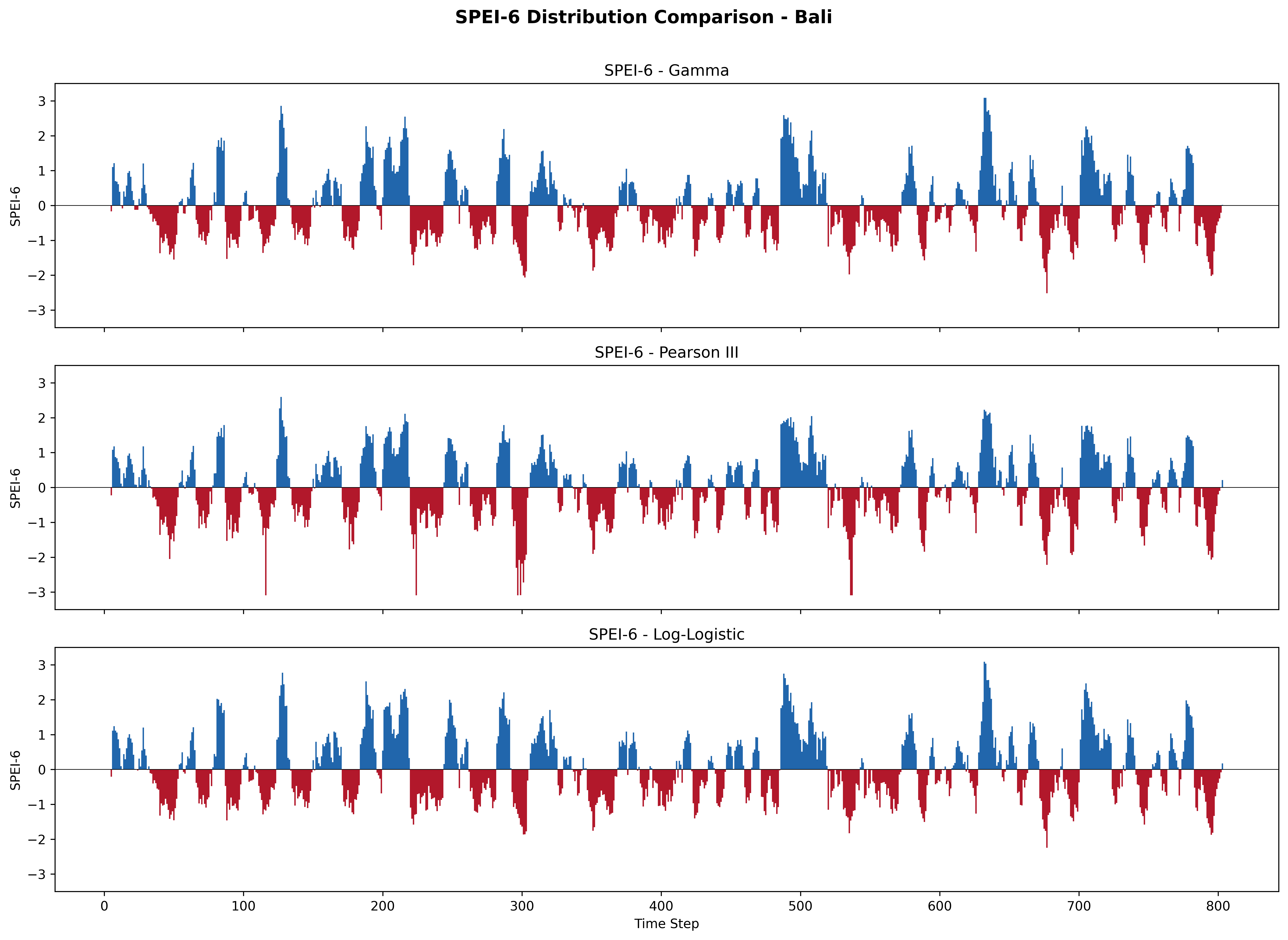
plt.suptitle(f'SPEI-6 Distribution Comparison - Bali', fontsize=14, fontweight='bold', y=1.01)
plt.tight_layout()
plt.show()
```
::: {.callout-tip}
### Distribution Recommendations for SPEI
- **Gamma** (default): Works well for most cases and is the fastest.
- **Pearson III**: Handles skewed water balance (P - PET) data well. Recommended when your region has highly seasonal PET.
- **Log-Logistic**: Provides better tail behavior for extreme events and is used in the original R SPEI package.
All three distributions produce highly correlated results (r > 0.98). The choice matters most for extreme event characterization.
:::
## 9. Spatial Analysis
Visualize SPEI-12 spatial patterns across Bali at different time periods.
```{python}
fig, axes = plt.subplots(2, 2, figsize=(14, 10))
axes = axes.flatten()
# Select recent time points
time_indices = [-48, -36, -24, -12] # 4, 3, 2, 1 years ago
titles = ['4 years ago', '3 years ago', '2 years ago', '1 year ago']
for i, (t_idx, title) in enumerate(zip(time_indices, titles)):
time_str = str(spei_12.time[t_idx].values)[:7] # Format: YYYY-MM
im = axes[i].pcolormesh(
spei_12.lon,
spei_12.lat,
spei_12[t_idx, :, :],
cmap='RdYlBu',
vmin=-2.5,
vmax=2.5,
shading='auto'
)
axes[i].set_title(f'{title} ({time_str})')
axes[i].set_xlabel('Longitude')
axes[i].set_ylabel('Latitude')
axes[i].set_aspect('equal')
cbar = plt.colorbar(im, ax=axes[i])
cbar.set_label('SPEI-12')
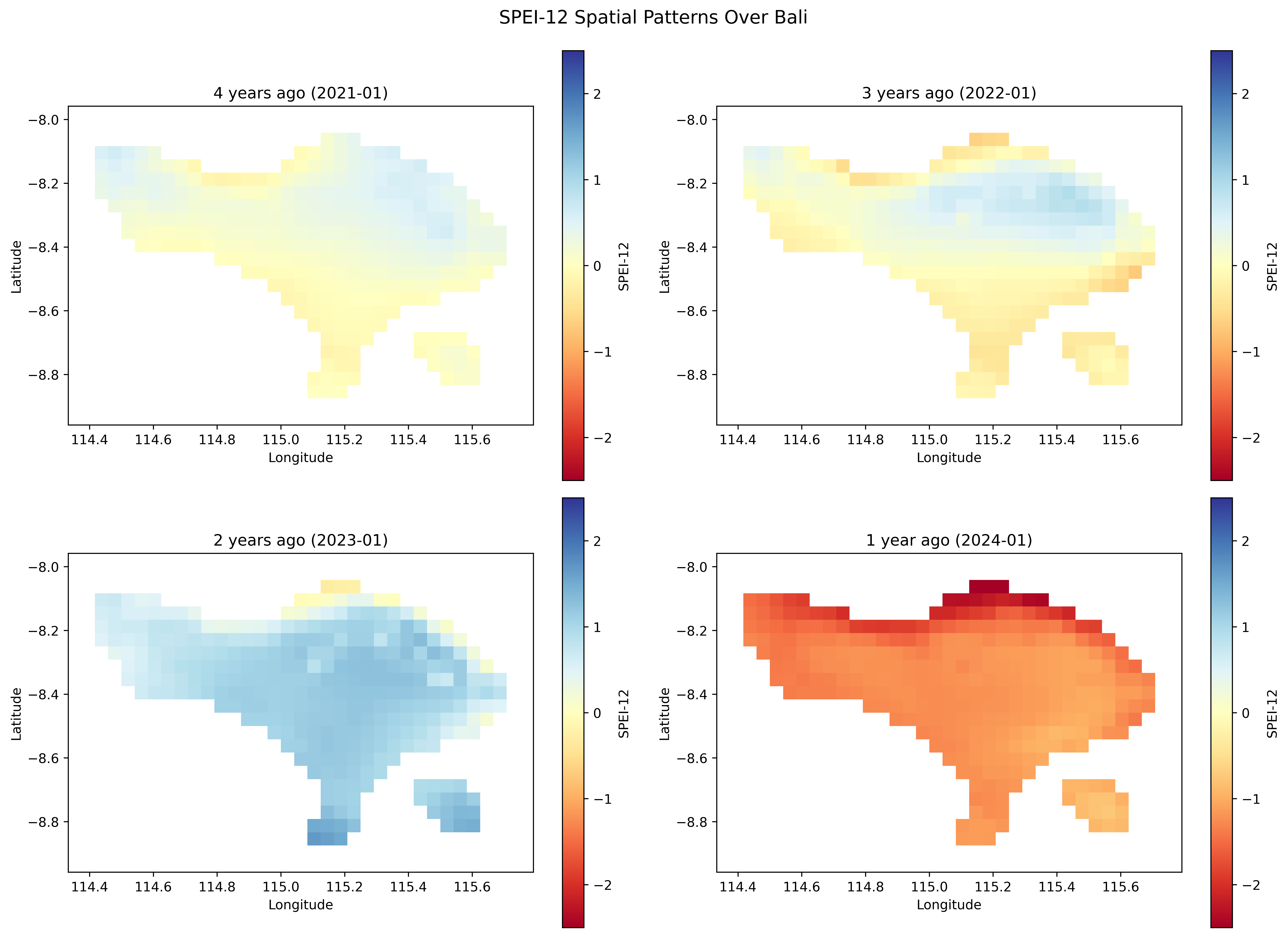
plt.suptitle('SPEI-12 Spatial Patterns Over Bali', fontsize=14, y=0.995)
plt.tight_layout()
plt.show()
```
The spatial maps show how drought conditions vary across Bali at different time periods. The RdYlBu color scale maps red to dry conditions and blue to wet conditions. Ocean cells appear as white (no data).
## 10. Drought Area Percentage Comparison
Compare the percentage of area experiencing drought conditions for SPI vs SPEI.
```{python}
# Calculate drought area for both SPI and SPEI
print("Calculating drought area percentages...")
spi_drought_area = get_drought_area_percentage(spi_12, threshold=-1.0)
spei_drought_area = get_drought_area_percentage(spei_12, threshold=-1.0)
print("Drought area calculation complete!")
```
```{python}
fig, ax = plt.subplots(figsize=(14, 5))
spi_drought_area.plot(ax=ax, linewidth=1.2, color='steelblue', label='SPI-12', alpha=0.7)
spei_drought_area.plot(ax=ax, linewidth=1.2, color='darkgreen', label='SPEI-12', alpha=0.7)
spi_mean = float(spi_drought_area.mean())
spei_mean = float(spei_drought_area.mean())
ax.axhline(y=spi_mean, color='steelblue', linestyle='--',
linewidth=0.8, alpha=0.5, label=f'SPI mean: {spi_mean:.1f}%')
ax.axhline(y=spei_mean, color='darkgreen', linestyle='--',
linewidth=0.8, alpha=0.5, label=f'SPEI mean: {spei_mean:.1f}%')
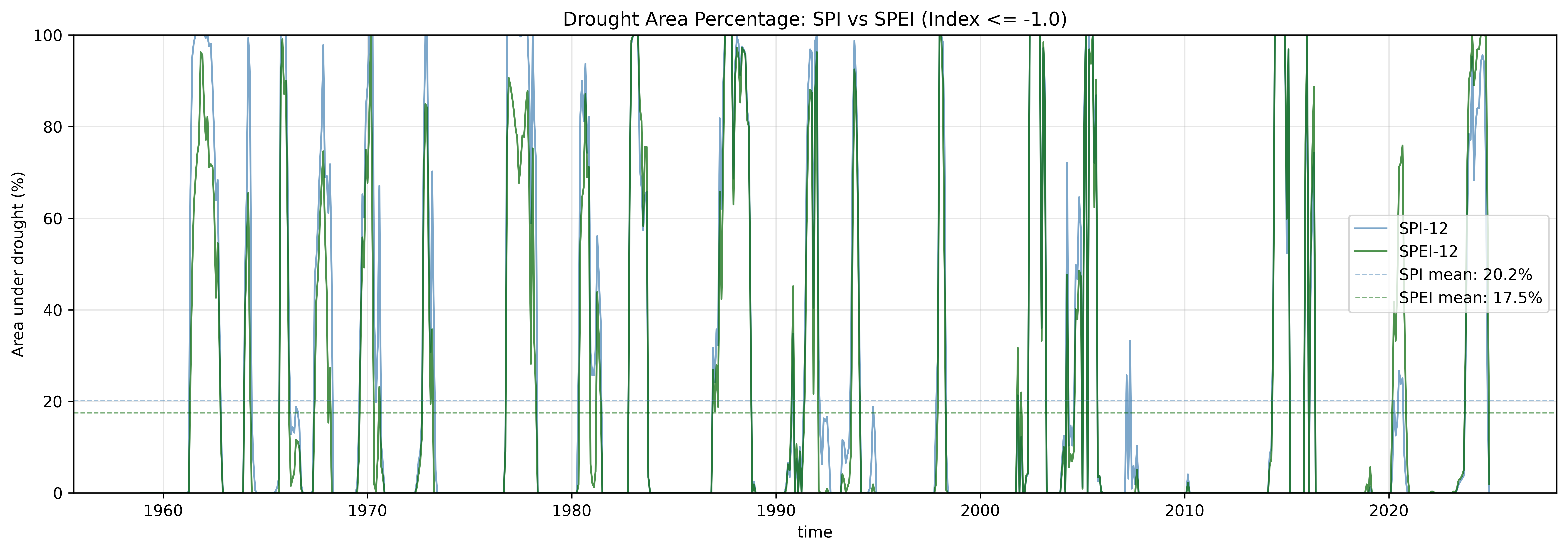
ax.set_title('Drought Area Percentage: SPI vs SPEI (Index <= -1.0)', fontsize=12)
ax.set_ylabel('Area under drought (%)')
ax.set_ylim(0, 100)
ax.grid(True, alpha=0.3)
ax.legend()
plt.tight_layout()
plt.show()
print(f"\nDrought Area Statistics:")
print(f" SPI-12 mean: {spi_mean:.1f}%")
print(f" SPEI-12 mean: {spei_mean:.1f}%")
print(f" Difference: {(spei_mean - spi_mean):.1f}%")
```
This comparison reveals how temperature/PET inclusion affects the drought signal. A higher mean drought area for SPEI suggests that evaporative demand is amplifying drought conditions beyond what precipitation alone would indicate.
## 11. Save and Reuse Fitting Parameters
```{python}
# Save SPEI fitting parameters for future reuse
params_file = str(output_netcdf / 'spei_gamma_params_12_month_bali.nc')
save_fitting_params(
params,
params_file,
scale=12,
periodicity='monthly',
index_type='spei',
calibration_start_year=calib_start,
calibration_end_year=calib_end,
coords={'lat': precip.lat, 'lon': precip.lon},
distribution='gamma'
)
print(f"Parameters saved to: {params_file}")
# Load parameters
loaded_params = load_fitting_params(params_file, scale=12, periodicity='monthly')
print(f"Parameters loaded successfully")
print(f" Alpha shape: {loaded_params['alpha'].shape}")
print(f" Beta shape: {loaded_params['beta'].shape}")
print(f" Prob_zero shape: {loaded_params['prob_zero'].shape}")
```
## 12. Save Results to NetCDF
```{python}
# Save SPEI-12
spei_12_file = str(output_netcdf / 'spei_12_bali.nc')
save_index_to_netcdf(spei_12, spei_12_file, compress=True, complevel=5)
print(f"SPEI-12 saved to: {spei_12_file}")
# Save multi-scale SPEI
spei_multi_file = str(output_netcdf / 'spei_multi_scale_bali.nc')
save_index_to_netcdf(spei_multi, spei_multi_file, compress=True, complevel=5)
print(f"Multi-scale SPEI saved to: {spei_multi_file}")
# If PET was calculated from temperature, save it for future use
if not use_precomputed_pet:
pet_file_out = str(output_netcdf / 'pet_thornthwaite_bali.nc')
pet_calculated = calculate_pet(temperature, precip.lat.values, data_start_year)
pet_calculated.to_netcdf(pet_file_out)
print(f"Calculated PET saved to: {pet_file_out}")
print("\nAll results saved successfully!")
```
## 13. Summary
### Key Takeaways
1. **PET Input Options**: Use pre-computed PET for accuracy, or temperature for convenience
2. **SPI vs SPEI**: SPEI captures temperature-driven drought that SPI misses
3. **Time Scales**: Choose based on application (1=meteorological, 3=agricultural, 6=hydrological, 12=water resources)
4. **Distribution Choice**: Gamma (default), Pearson III, or Log-Logistic --- all produce consistent results (r > 0.98)
5. **Climate Change**: SPEI is essential for regions with warming trends
6. **Spatial Analysis**: Drought conditions vary across the landscape
7. **Parameter Reuse**: Save fitting parameters for faster operational processing
### Recommended Applications
| Application | Recommended Scale |
|:---|:---|
| Crop drought monitoring | SPEI-3 or SPEI-6 |
| Forest fire risk | SPEI-6 or SPEI-12 |
| Water resources management | SPEI-12 or longer |
| Climate change studies | Compare SPI vs SPEI trends |
| Vegetation stress | SPEI-3 or SPEI-6 |
## Next Steps
- **[Event Analysis](03-event-analysis.qmd)** - Identify and characterize drought events using SPEI
- **[Visualization Gallery](04-visualization.qmd)** - Create publication-quality plots
## References
- McKee et al. (1993): SPI methodology
- Vicente-Serrano et al. (2010): SPEI methodology
- Thornthwaite (1948): PET calculation (Thornthwaite method)
- Hargreaves & Samani (1985): PET calculation (Hargreaves method)